By A Mystery Man Writer

How to convert the molecular graph in SDF/JSON format to the input of graphormer? · Issue #108 · microsoft/Graphormer · GitHub

Relating the xyz files to the molecules in PCQM4Mv2 · Issue #297 · snap-stanford/ogb · GitHub
sdftosmi.py: Convert multiple ligands/compounds in SDF format to SMILES. — Bioinformatics Review
Pytorch-Geometric/pytorch_geometric_introduction.py at master · marcin-laskowski/Pytorch-Geometric · GitHub

Metapath-fused heterogeneous graph network for molecular property prediction - ScienceDirect
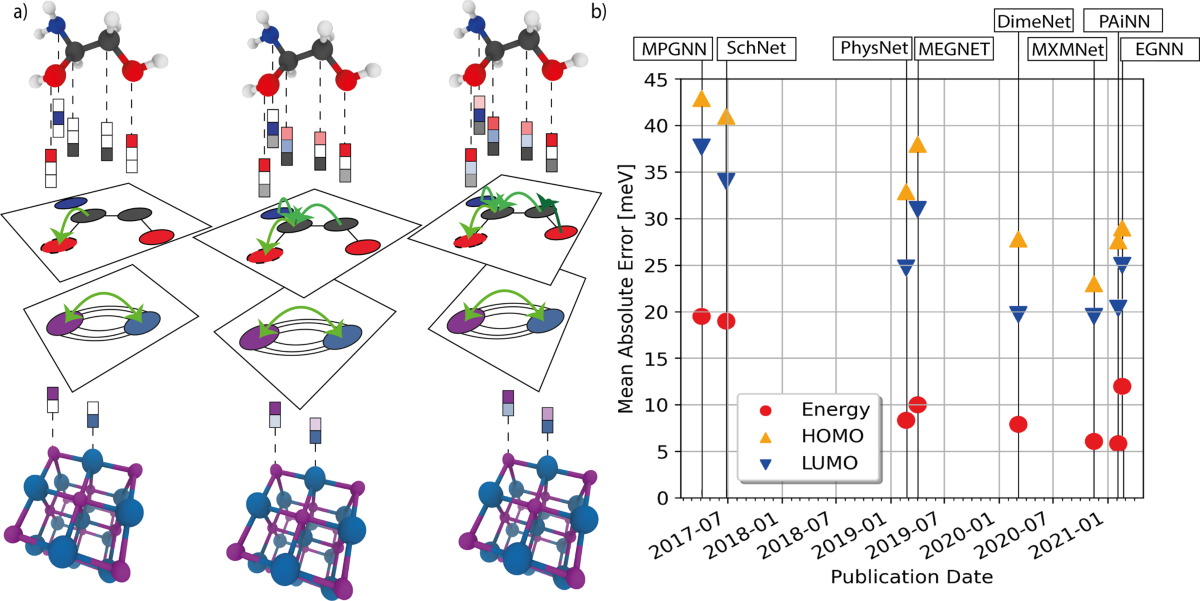
Graph neural networks for materials science and chemistry

Hands-On Graph Neural Networks Using Python Practical Techniques and Architectures For Building Powerful Graph and Deep Learning Apps With PyTorch PDF, PDF, Machine Learning

PDF) Hierarchical Grammar-Induced Geometry for Data-Efficient Molecular Property Prediction
Data — kgcnn 2.2.1 documentation
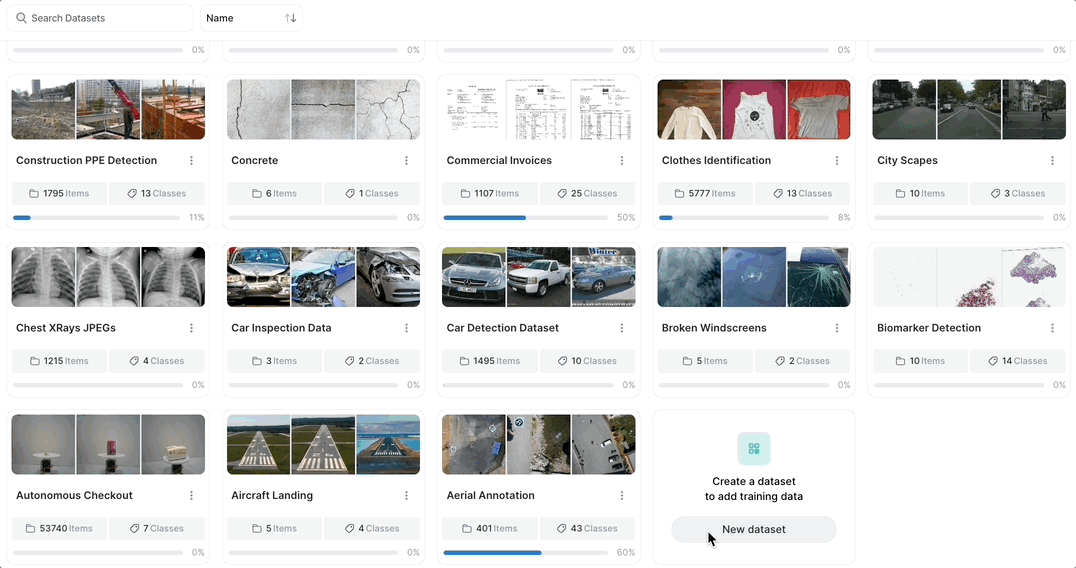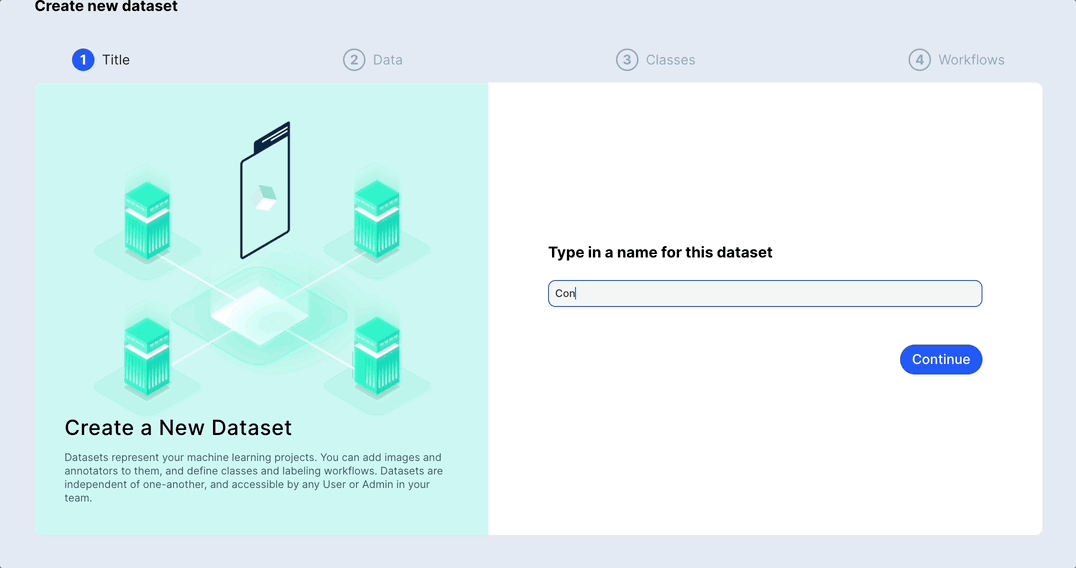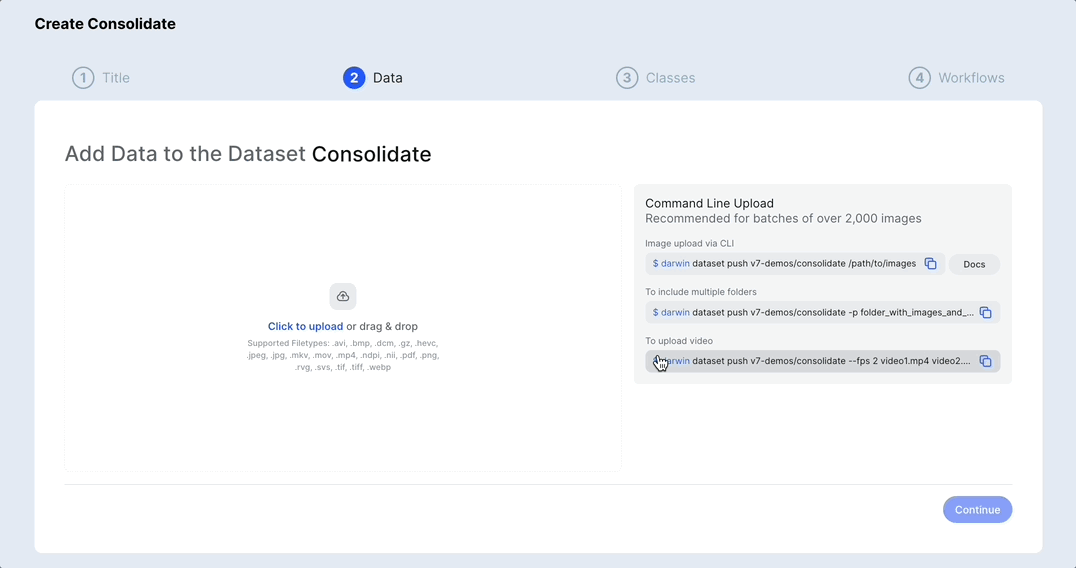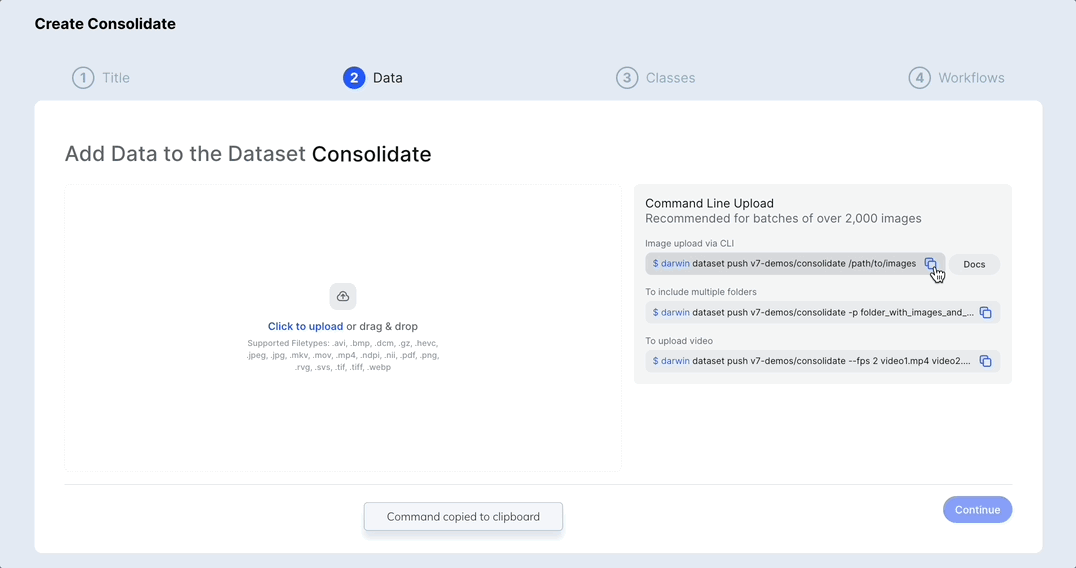
Merge or duplicate datasets
File Format
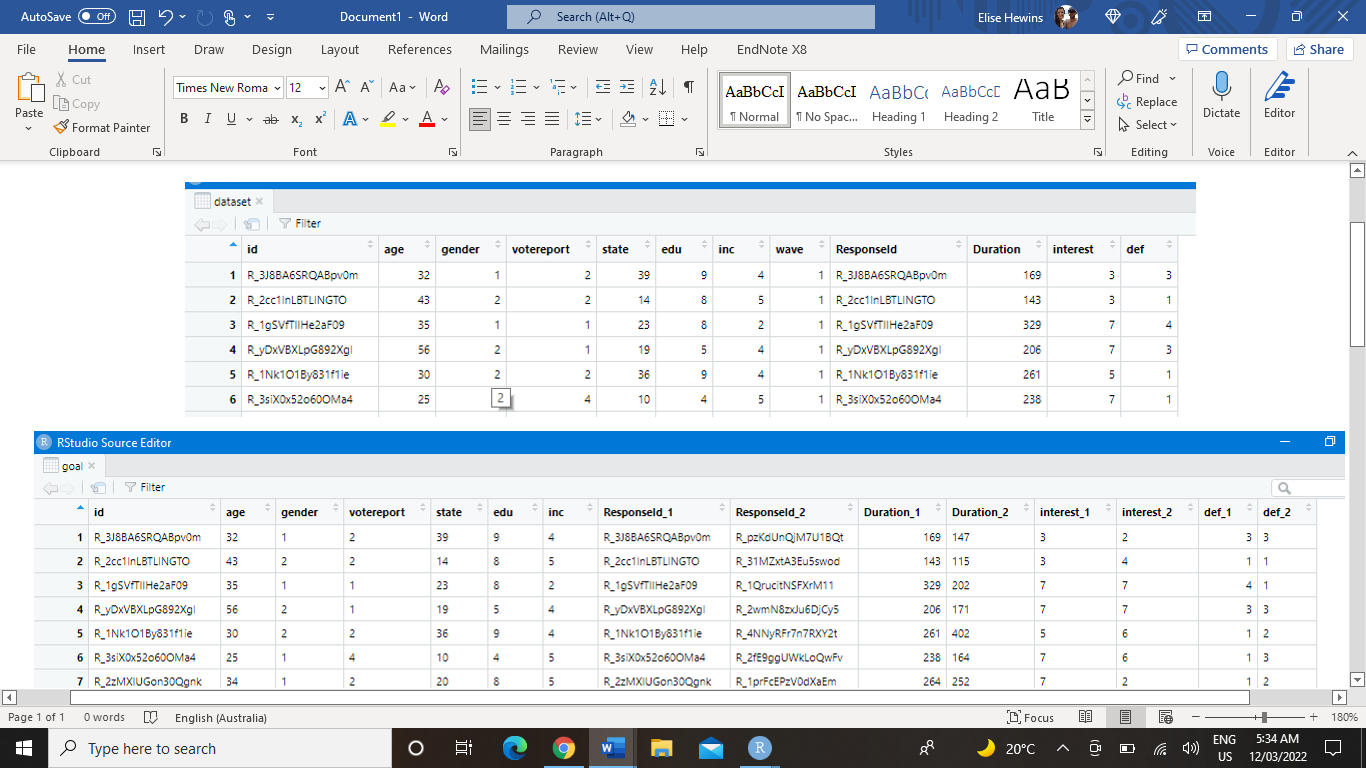
converting a dataset from a long to wide format. - tidyverse - Posit Community